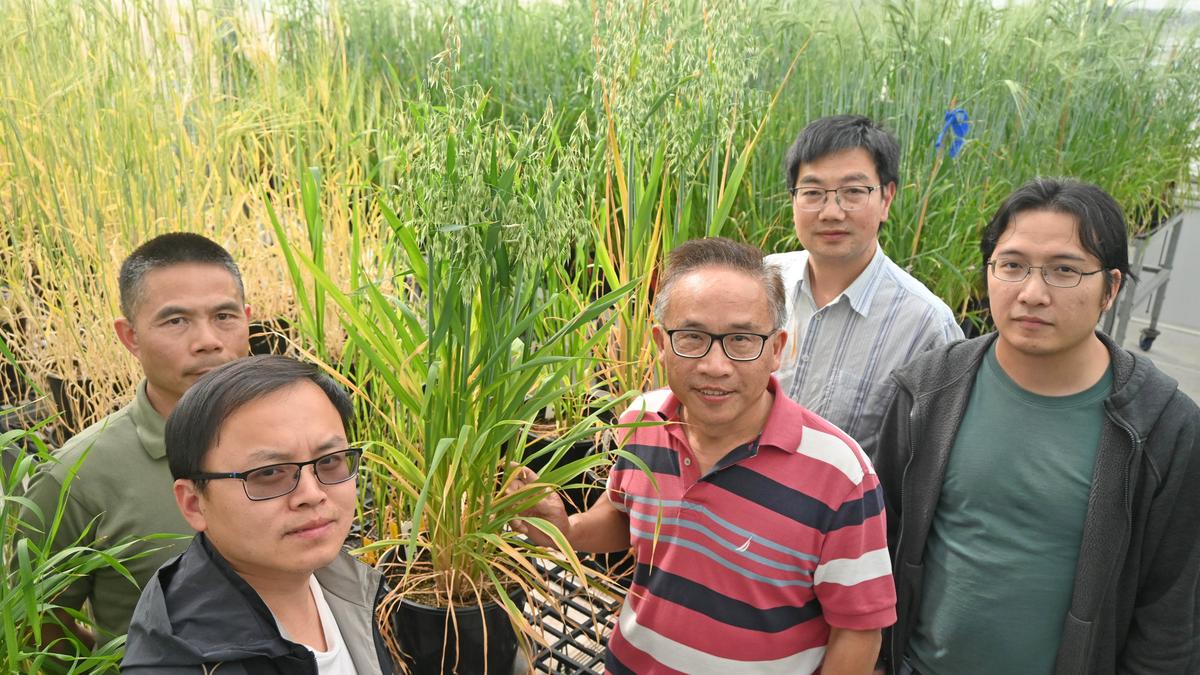
A significant international collaboration has successfully decoded the pangenome of oats, a cereal known for its genetic complexity and nutritional value. Researchers from Murdoch University and the Department of Primary Industries and Regional Development in Western Australia contributed vital genome sequences from local oat varieties, including Bannister, Bilby, and Williams, to this groundbreaking study published in the journal Nature.
This research effort was spearheaded by the IPK Leibniz Institute in Germany and involved over 70 scientists from 33 institutions across 10 countries. The resulting findings provide an invaluable blueprint for understanding oat genetic diversity, pinpointing genes associated with yield, plant health, and resilience to environmental changes. Such insights could revolutionize oat breeding practices and crop improvement initiatives globally.
Professor Chengdao Li, director of the Western Crop Genetics Alliance and leader of the research team at Murdoch University, emphasized the importance of this study. He stated, “This research transforms oats from a genetic “black box” into a blueprint that will enable precision breeding for a healthier, more sustainable food future.” The identification of specific genetic markers, such as the 2A/2C gene translocation found in Australian oats, illustrates the natural adaptation of crops to their environments.
By leveraging this knowledge, breeders can develop more resilient oat varieties tailored to specific regional conditions. The study sequenced and analyzed 33 oat lines, incorporating both cultivated varieties and their wild relatives, utilizing advanced genomic and transcriptomic technologies. A unique “pantranscriptome” was also established, detailing gene activity across various tissues and developmental stages.
Despite the loss of numerous genes in one of the crop”s three subgenomes, oats maintain impressive productivity due to other gene copies compensating for these deficiencies. Additionally, the research uncovered that structural rearrangements in oat DNA—such as inversions and translocations—have played a critical role in the crop”s adaptation and domestication processes.
Dr. Kaara Klepper, executive director of broadacre systems at DPIRD, remarked on the significance of these findings, stating, “The decoded oat pangenome epitomizes how genomics research is fueling advances in crop breeding, agricultural production, and human health.” He highlighted the substantial contributions made by scientists from DPIRD and Murdoch University, which will aid Western Australian farmers in cultivating high-performance, resilient crops that are better suited to a changing climate, enhancing both sustainability and profitability.
Known for their health advantages—including high fiber content, cholesterol-lowering properties, and gluten-free characteristics—oats have long posed a challenge for researchers due to their complex genome, composed of six sets of chromosomes from three ancestral species. With the oat pangenome now decoded, scientists are better equipped to enhance yield, nutritional value, and climate resilience, heralding a new era in oat breeding and sustainable agricultural practices.