Researchers at the University of Tokyo have unveiled an innovative method for analyzing proteins that combines machine learning with voltage-matrix nanopore profiling. This technique addresses the challenges faced by existing analytical tools in distinguishing subtle structural differences among biomolecules, a crucial aspect in biomedical research and diagnostics.
The study, published in Chemical Science, introduces a method that utilizes multivoltage solid-state nanopore recordings to classify proteins based on their intrinsic electrical signatures. This approach allows for the identification and categorization of “molecular individuality” without the need for labeling or modifications.
Solid-state nanopores are tiny openings through which proteins or other molecules can pass, facilitated by ionic current. By manipulating voltage during this process, the signals generated as proteins traverse the nanopores can assist in identifying the molecules. Previous applications of nanopore technologies have primarily focused on DNA and RNA analysis, with limitations in protein analysis due to their complex structures and variable signal behaviors.
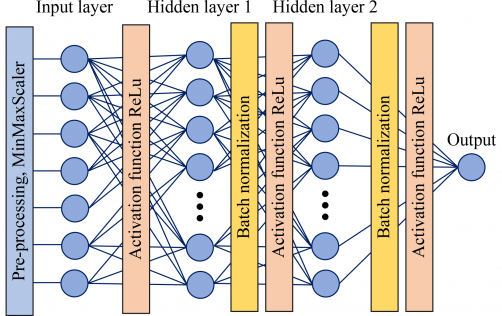
The research team”s novel strategy involves systematically altering voltage conditions to capture both stable and voltage-dependent signal patterns. By organizing these data into a voltage matrix, the machine learning model can successfully differentiate proteins even within complex mixtures, expanding nanopore applications beyond sequencing to general molecular profiling.
Professor Sotaro Uemura, from the Department of Biological Sciences, commented on the difficulty of identifying and classifying proteins in complex biological samples. Traditional techniques like enzyme-linked immunosorbent assay (ELISA) and mass spectrometry often struggle to identify subtle structural variations without the use of labels. He noted that while solid-state nanopores present a promising solution, earlier methods were constrained by their dependence on single-voltage measurements. Their research aimed to overcome this limitation.
To validate their approach, the researchers analyzed mixtures containing two cancer-related protein biomarkers: carcinoembryonic antigen (CEA) and cancer antigen 15-3 (CA15-3). By constructing a voltage matrix from signals recorded at six different voltage levels, they successfully identified distinct response patterns for each protein. Additionally, the method detected changes in molecular populations when an aptamer, a synthetic DNA segment, was attached to CEA.
To further demonstrate the method”s practicality, the team applied the voltage-matrix framework to mouse serum samples. By comparing samples that had undergone centrifugation with those that had not, the researchers found that the two sample types could be clearly differentiated within the voltage matrix. This finding suggests that their method can identify and categorize subtle compositional differences in complex biological samples, indicating its potential for real-world bioanalytical applications.
Uemura stated, “By systematically varying voltage conditions and employing machine learning, we create a voltage matrix that reveals both robust, voltage-independent molecular characteristics and voltage-sensitive structural alterations.” He emphasized that their study establishes a novel approach to represent and classify molecular signals across varying voltages, facilitating the visualization of molecular individuality and estimation of compositions within mixtures.
Looking to the future, the research team intends to expand their framework to include human serum or saliva samples. They also plan to develop a parallelized nanopore system capable of performing multiple tasks simultaneously, enabling real-time molecular profiling. This foundational work could pave the way for applications ranging from biomedical diagnostics to environmental monitoring.