A research team has identified and characterized 15 abscisic acid (ABA) receptor genes in tomatoes, with a focus on two specific receptors, SlPYR1.1 and SlPYL8.1, which play crucial roles in balancing growth and defense mechanisms. Overexpression of these receptors led to increased sensitivity to ABA, resulting in dwarfism, delayed aging, reproductive issues, and enhanced fruit pigmentation.
The study emphasizes the importance of ABA signaling in regulating hormonal interactions that fine-tune tomato growth, aging, and reproduction. While ABA is known for its essential role in plant stress responses and fruit ripening, its influence on vegetative growth and flower development has not been thoroughly explored.
ABA signaling operates through the PYR/PYL/RCAR receptor family, which acts as molecular switches that activate downstream kinases to initiate physiological responses. Although these receptors have been studied in model plants like Arabidopsis, the varied and overlapping functions of receptor subfamilies have complicated the understanding of how ABA impacts developmental flexibility. Tomatoes serve as an ideal model for studying fruit development and hormonal interactions, offering insights into these processes.
The research, published in Plant Hormones on July 8, 2025, was conducted by a team from Chongqing University of Arts and Sciences. The researchers utilized tomato genome databases and transcriptomic datasets to identify members of the PYR/PYL/RCAR receptor family. The 15 identified SlPYR/PYL genes were classified into three subfamilies based on their Arabidopsis homologs.
Expression profiling revealed that SlPYR1.1 and SlPYL8.1 were significantly responsive to ABA, salt, and ethylene, with the highest expression levels found in roots and leaves, although their expression was suppressed during fruit ripening. Subcellular localization studies in Nicotiana benthamiana demonstrated that these proteins reside in both the nucleus and cytoplasm, suggesting a dual role in regulating ABA-mediated transcription.
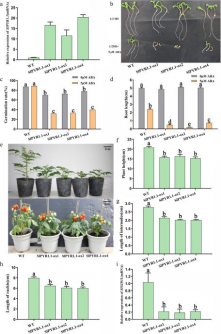
Functional validation through overexpression lines showed that plants with elevated SlPYR1.1 levels exhibited increased ABA sensitivity, lower germination rates, shorter roots, and pronounced dwarfism due to the downregulation of SlGID1, a vital gibberellin receptor. This finding indicates a competitive relationship between ABA and gibberellin pathways.
Conversely, SlPYL8.1 overexpression resulted in delayed leaf aging, characterized by dark green, thickened leaves with higher chlorophyll and carotenoid content due to enhanced mesophyll cell proliferation. However, this overexpression also caused reproductive anomalies, such as floral bud abortion and altered inflorescence orientation, highlighting developmental trade-offs.
Notably, fruits from SlPYL8.1 overexpression plants displayed increased lycopene accumulation and a deeper red color, suggesting that this receptor not only regulates stress and growth but also plays a role in fruit coloration and ripening. The findings underscore the importance of SlPYR1.1 and SlPYL8.1 in linking ABA signaling with developmental and stress-related responses.
Adjusting the expression of these receptors could pave the way for achieving desirable agricultural traits, including improved stress resilience, delayed leaf aging, and enhanced fruit color, without compromising overall plant health. The ability of SlPYL8.1 to boost lycopene levels presents a promising strategy for enhancing the nutritional quality and shelf life of tomatoes. Additionally, understanding how these receptors inhibit reproduction can inform breeding strategies aimed at minimizing yield losses.
The study was supported by the Chongqing talent program and the Chongqing Natural Science Foundation Innovation and Development Joint Fund Project.
Plant Hormones is an open-access journal that publishes peer-reviewed research on plant hormone biosynthesis, signal transduction, and the implications for agriculture, horticulture, and forestry.